Soil microbial diversity and community composition of grassland and forest ecosystems along land-use gradients
Living organisms that are smaller than 50µm are classified as microorganisms. These include bacteria, archaea, fungi, but also unicellular smaller eukaryotes such as protists.
Microorganisms (i.e. Archaea, Bacteria and Fungi) represent the dominant life forms in soil both in terms of biomass and biodiversity and build complex functional interaction networks. These microbial communities are therefore often referred to as “soil microbiome”. They provide important ecosystem services, such as soil formation, element cycling, plant nutrition, detoxification, as well as the exchange of gas and matter with atmosphere and ground water. These key roles rely on complex trans-kingdom interactions between the soil microorganisms themselves but also between microorganisms and the below and aboveground plant and animal communities.
In addition to site-specific properties such as soil type and local climate, the types of land use and land use intensity can also be important drivers of soil microbial diversity. So far, the functional relationships between environmental factors and microbial diversity in soil are not sufficiently understood.
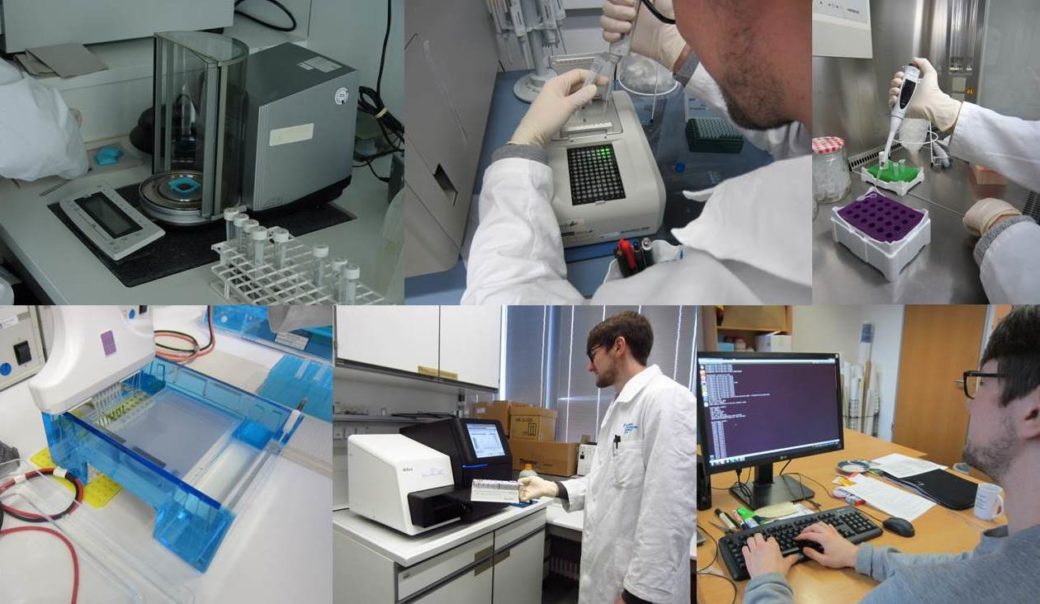
Thanks to the development of high throughput sequencing and powerful bioinformatics methods, it has become possible to analyse the biodiversity of soil microorganisms at large scales. Since the start of the Biodiversity Exploratories in 2006, Core project 8 has been in charge of monitoring the biodiversity of soil microorganisms.
While the initial focus was on soil and arbuscular mycorrhizal fungi, Core 8 in its current phase encompasses the study of all relevant microorganisms, including fungi, bacteria and archaea.
The main objectives of the Microorganisms Core Project are:
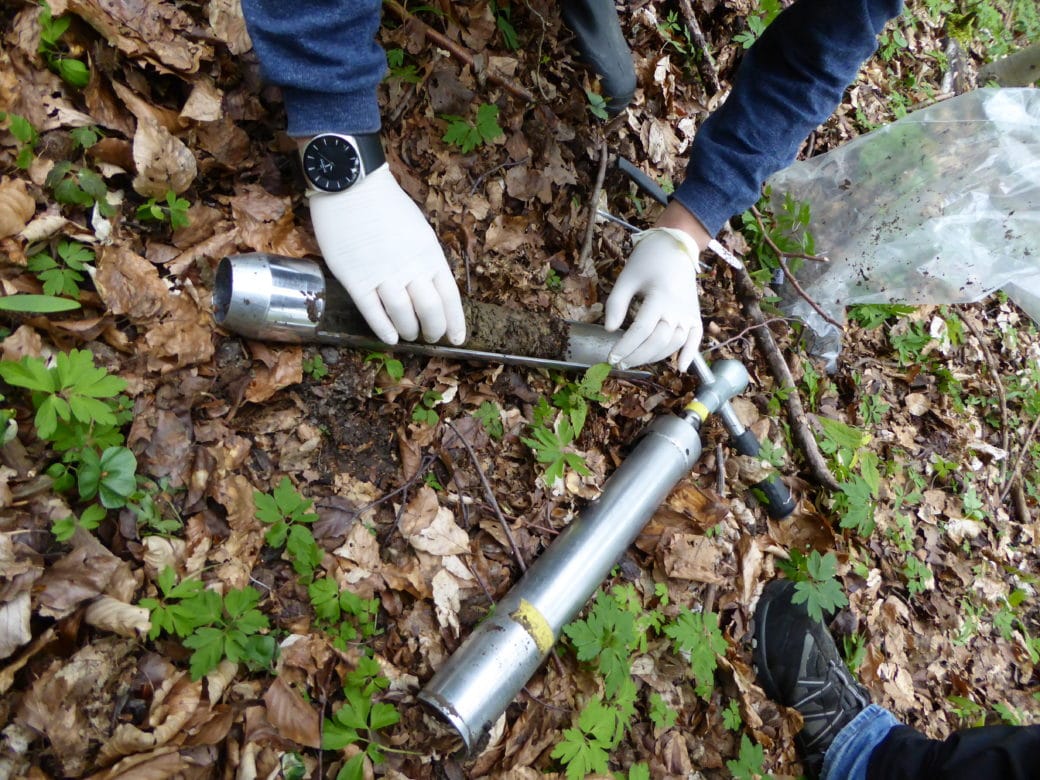
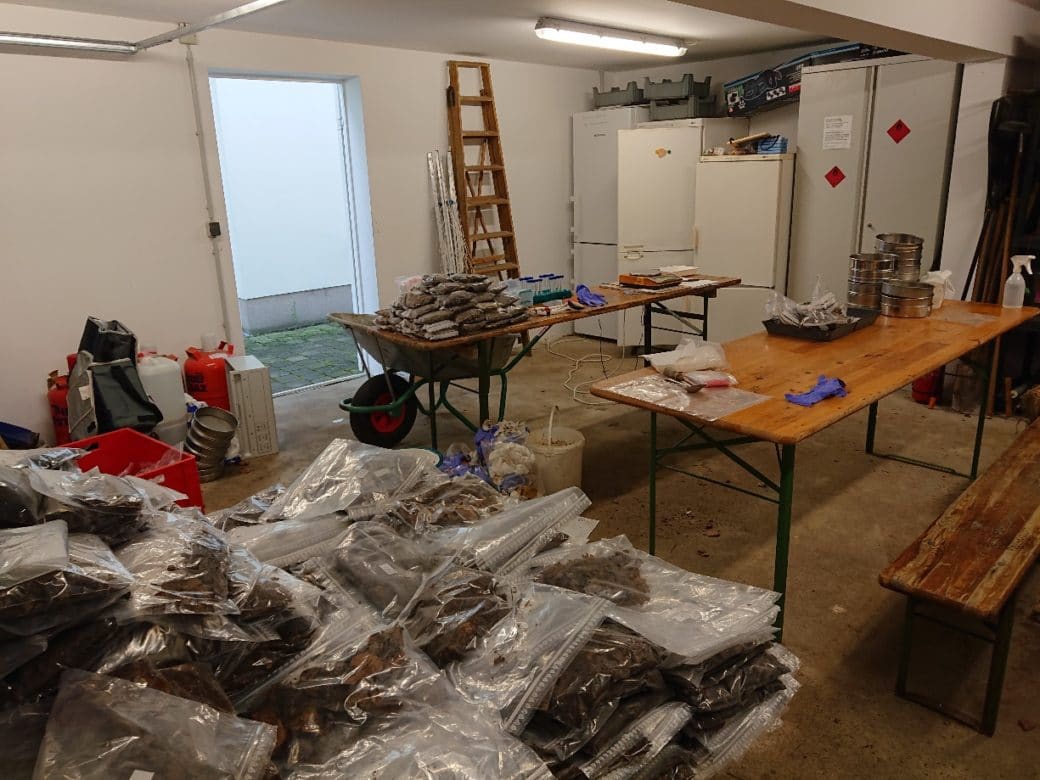
- Organization of coordinated soil sampling campaigns (in cooperation with Core project 9). The next campaign is planned for spring 2023.
- Long-term archive of soil samples und extracted nucleic acids.
- Long term monitoring on biodiversity of soil bacteria, archaea and fungi in all 300 EPs as well as in the newly established field experiments (FOX, REX I+II, LUX).
- Identification and characterization of microbial key stone species employing co-occurrence network analyses, followed by selected metagenomics analyses using shot gun sequencing. This will provide better insights into functional microbial diversity.
- Identification and assessment of the small but functionally relevant proportion of active bacteria, through analysis of rRNA/rDNA ratios. This is of relevance, since the majority of soil bacteria is inactive or dormant and hence does not contribute to ecosystem functions.
- Establishment of adequate primers and quantification of key organisms using quantitative PCR (qPCR).
- Functional fingerprinting and reconstruction of microbial genomes from representative grassland areas (metagenomes).
- Participation in several synthesis activities within the Biodiversity Exploratories community.
- Interaction and data exchange with projects on an international level.
All core projects provide important basic information on land use, diversity and ecosystem processes (long-term monitoring). These are made available to the sub-projects in each phase for researching more in-depth questions.
Services provided in the current phase
In the 6th phase (2020-2023) the core project 8 provides the following services / basic examinations:
- Organization, in cooperation with Core project 9, of the coordinated soil sampling campaigns of 2021 (under Corona restrictions) and 2023.
- Deposit and long-term storage (-80°C) of soil samples from coordinated soil sampling campaigns and new experiments (FOX, REX I+II, LUX) for subsequent analysis. Subsamples can be provided for future work in the contributing projects (targeted at molecular work – small quantities).
- Standardization of nucleic acid extraction techniques as well as up and downstream bioinformatics after next generation sequencing. Optimized standards can be shared with contributing projects.
- Extraction and storage of nucleic acids (both DNA and RNA) from all EP plot samples from coordinated soil sampling campaigns and new experiments (FOX, REX I+II, LUX), which can be provided to demanding contributing projects.
- Provision of diversity data on soil microorganisms (Archaea, Bacteria and Fungi).
- Sequencing (Illumina, PacBio) for contributing projects.
- Training and assistance to members of contributing projects on molecular techniques and bioinformatics after next generation sequencing.
Services provided in past phases
- Provision of diversity data on soil Fungi. We already uploaded lists of amplicon sequence variants (ASVs) or operational taxonomic units (OTUs) of soil and arbuscular mycorrhizal fungi for the sampling campaigns 2011, 2014 and 2017 in all EPs. This data has been used in numerous synthesis publications on the relationship between land use intensity, below and above ground biodiversity and multi-functionality of ecosystems.
- Provision of DNA extracts from soil samples of coordinated soil sampling campaigns (2008-2017) to all interested contribution projects.
- Quantification of key organisms using quantitative PCR (qPCR) (2008-2017).
Core 8 data were used to:
- analyse the distance decay of forest soil microbial communities across Germany and how tree species modulate it in the soil rooting zone (Goldmann et al. 2016. Scientific reports, 6(1), 1-10).
- unravel the spatiotemporal variability of arbuscular mycorrhizal fungi grasslands (Goldmann et al. 2020. Environmental Microbiology, 22(3), 873-888).
- unravel the spatiotemporal variability of nitrifying organisms and their interactions (Stempfhuber et al. 2017).
- reveal that in grasslands, belowground biodiversity reacts opposite to aboveground biodiversity with increase of land use intensity (Goßner et al. 2016. Nature, 540(7632), 266-269).
- demonstrate that biodiversity at multiple levels is needed for ecosystem multifunctionality (Soliveres et al. 2016. Nature, 536(7617), 456-45) and rare species influence grassland multifunctionality (Soliveres et al. 2016. Nature, 536, 456-459).
- identify via rRNA/rDNA ratios and analyse the fraction of active soil bacteria and their contribution to soil function (in preparation).